Every year, thousands of life science students learn Python, bioinformatics tools, and different software. They complete online courses, collect certificates, and add skills to their resumes.
Then they apply for internships, research positions, or PhD programs and get rejected.
Not because they are not capable.
But because they cannot prove they can do real work.
This is where most students make a mistake. They focus too much on learning and not enough on building projects.
That is exactly why I wrote a complete beginner-friendly guide on a molecular docking project. Not just theory, not just software installation, but the full workflow from start to finish, including how to present the project in your resume, portfolio, and GitHub.
If you are from biotechnology, bioinformatics, microbiology, biochemistry, or any life science background, this type of project can literally change your profile.
You can read the full guide here:
https://btgenz.in/blog/molecular-docking-project-beginners-guide
Why Molecular Docking Is a Powerful Project for Students
Molecular docking sits at the intersection of three important fields:
Biology
Chemistry
Computational analysis
That means a single project can demonstrate multiple skills:
Understanding protein structure
Understanding drug molecules
Using computational tools
Analyzing scientific results
Presenting scientific findings
Very few student projects demonstrate all these skills together. That is why docking projects are highly valuable for portfolios and research applications.
But the problem is, most tutorials on the internet either:
Are too technical for beginners, or
Are too simplified and not useful for real projects
So students end up watching tutorials but never actually completing a proper project.
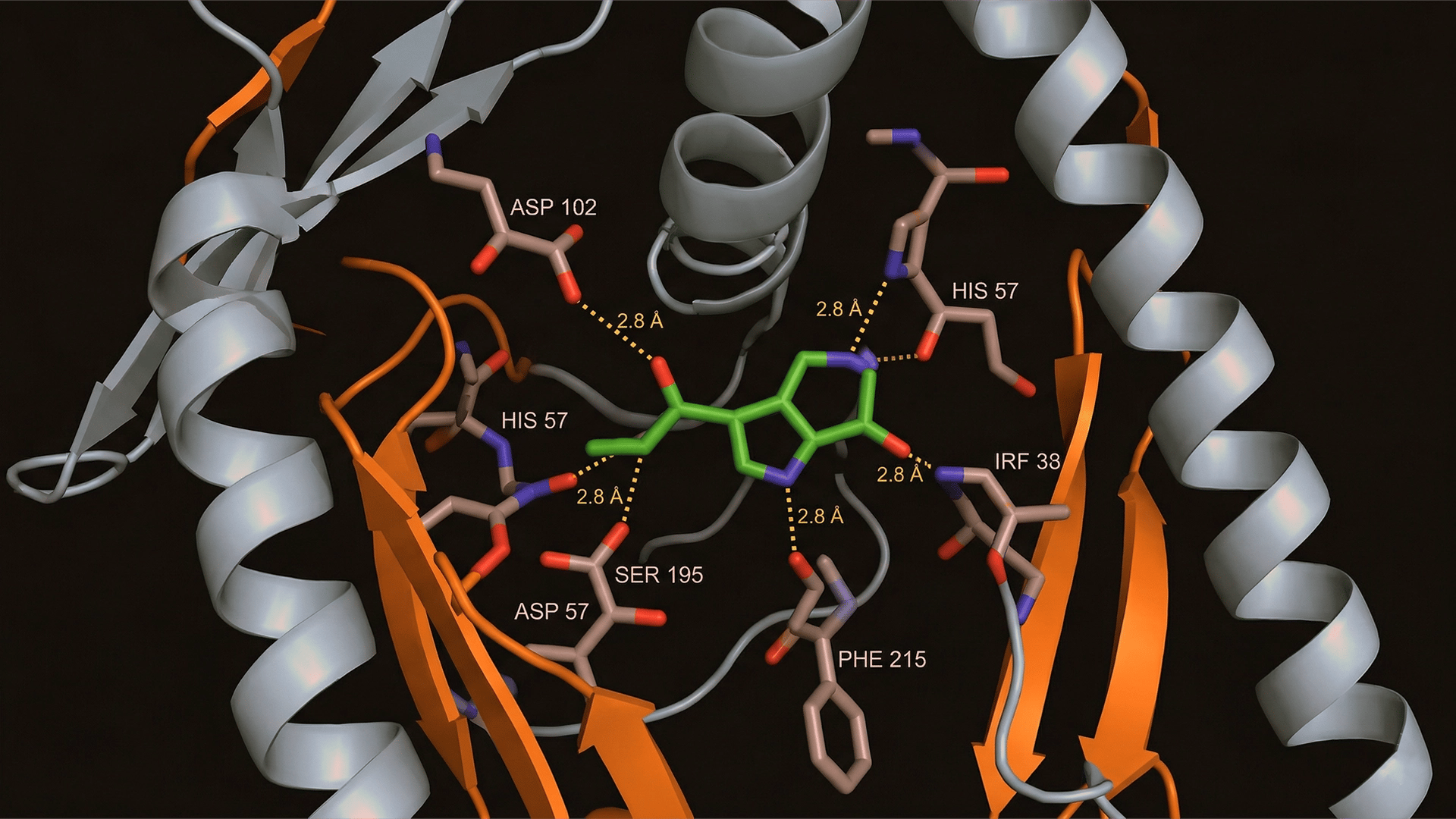
Protein Legend
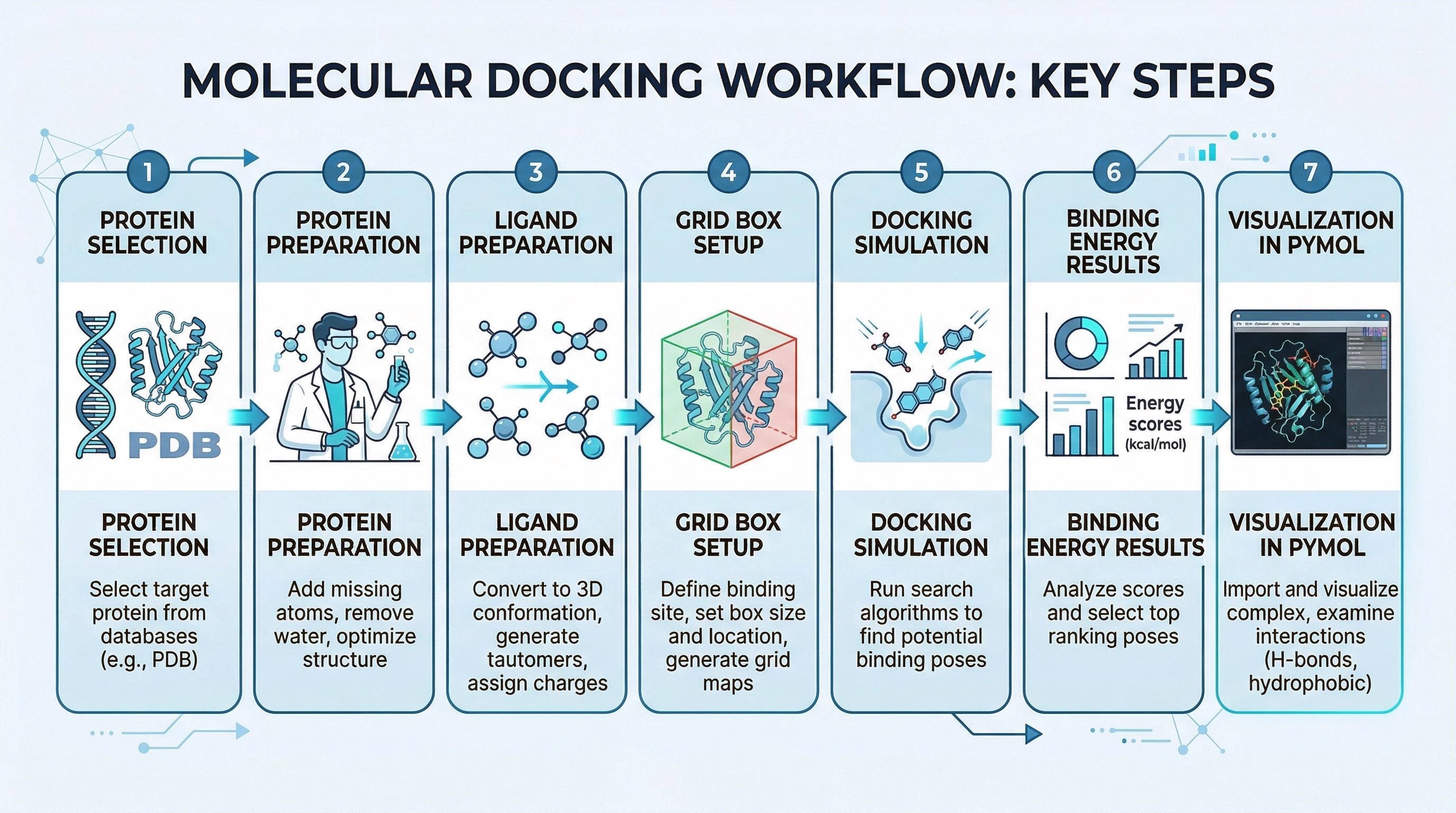
What This Guide Actually Teaches
The guide is designed like a real project, not like a software tutorial. It explains:
How to choose a target protein
Where to download protein structures
Where to download drug molecules
How to prepare protein and ligand files
How to define the binding site
How to run docking using AutoDock Vina
How to analyze binding energy
How to visualize protein-ligand interactions
How to present the project in a resume and a portfolio
What selection committees actually look for in such projects
So it is not just “how to run software”.
It is how to build a project that proves your skills.
If you want to read the full step-by-step guide, you can read it here:
https://btgenz.in/blog/molecular-docking-project-beginners-guide
Who Should Definitely Read This Guide
This guide is especially useful for:
- BSc Biotechnology students
- MSc Biotechnology students
- Bioinformatics students
- Life science students who want to move into computational biology
- Students preparing for research internships
- Students planning for a PhD abroad
- Students building a bioinformatics or computational biology portfolio
If you are trying to move from theory to practical work, this guide will help you understand how real computational biology projects are done.The Big Idea You Should Remember
Courses teach you concepts.
Projects prove you can work.
When selection committees or professors evaluate students, they don’t just ask:
What did you study?They ask:
What have you built?
What have you analyzed?
What problems have you solved?A molecular docking project answers all of these questions.
That is why this is not just a bioinformatics exercise.
This is a portfolio project.
Read the Full Guide
If you want to understand molecular docking step by step and also learn how to present it as a project for internships, research positions, or PhD applications, you can read the full guide here:
Read it slowly, and if possible, try to actually do the project while reading. That is where the real learning happens.